30 March 2026
Challenge accepted! Generating reference genomes and building a reference barcode library
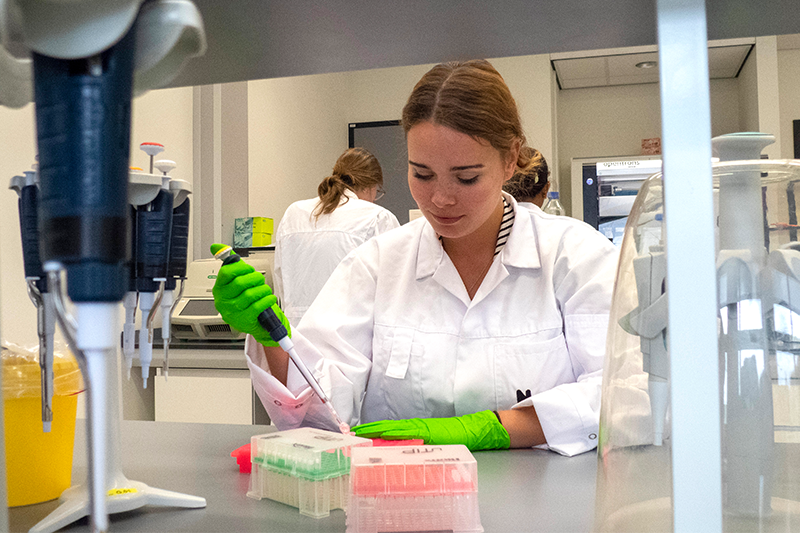
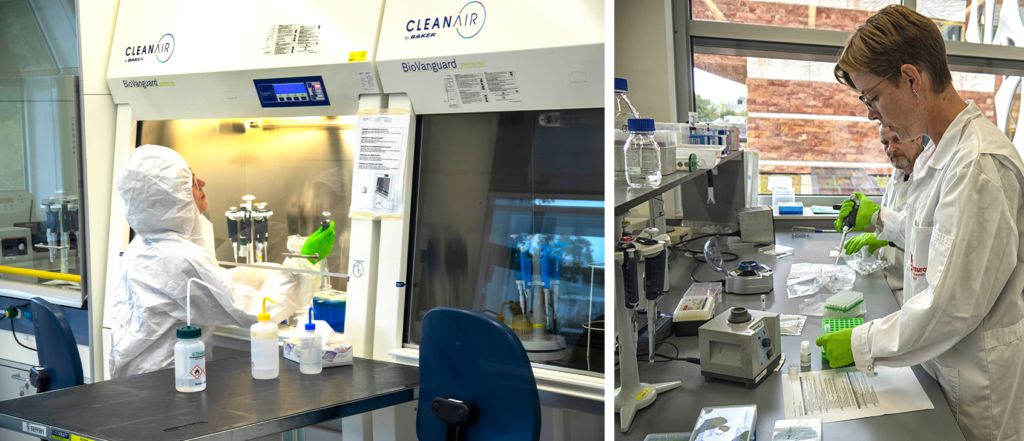
DNA sequencing covers a range of methods shaped by data needs, sample numbers, and sources. Within the BGE sequencing pillar, tens of thousands of DNA barcodes were generated from fresh and museum specimens, in addition to bulk collections and environmental samples of soil and seawater. Hundreds of whole-genome datasets were also produced, for which consistently higher-quality samples were required that posed fewer initial challenges, though the sequencing process itself was far more demanding. Key synergies were identified between DNA barcoding and genome sequencing on sharing expertise for challenging samples, negotiating cost-effective consumables and services, and aligning sequence data and metadata practices. These efforts improved progress tracking, and the integration and accessibility of BGE’s sequence data.
Want to know more about why fresh samples are so difficult to sequence successfully?
Read the interview with Olga below! ↓
From samples to reference genomes and building a reference barcode library: an interview with Olga Vinnere Pettersson
 To generate reference genomes and build a reference barcode library for European species, over 150,000 samples were collected from freshly caught specimens and museum specimens. These samples have all been processed in BGE’s consortium labs, but unfortunately not all 150,000 samples could be successfully sequenced. Although the library building of museum samples resulted in 90.6% success, the fresh samples proved challenging. Olga Vinnere Pettersson is one of the scientists dealing with sequencing the collected samples. She can tell us more about why fresh samples are so difficult to sequence successfully.
To generate reference genomes and build a reference barcode library for European species, over 150,000 samples were collected from freshly caught specimens and museum specimens. These samples have all been processed in BGE’s consortium labs, but unfortunately not all 150,000 samples could be successfully sequenced. Although the library building of museum samples resulted in 90.6% success, the fresh samples proved challenging. Olga Vinnere Pettersson is one of the scientists dealing with sequencing the collected samples. She can tell us more about why fresh samples are so difficult to sequence successfully.
“The biggest lesson from BGE’s sequencing work is deceptively simple,” Olga tells us. “Biodiversity is not just a scientific concept – it is a laboratory reality.” Standard genomics protocols are built around model organisms. They work beautifully for many species, but when you scale up to the full range of European biodiversity the exceptions multiply fast. “We encountered insects where standard extraction yielded too little or too fragmented DNA, plants whose massive repetitive genomes resisted routine library preparation, and marine invertebrates where tissue preservation and DNA quality did not go hand in hand,” she explains. “Across a relatively small species subsample sequenced within BGE, we flagged over 30 species that required significant protocol adaptation.”
According to Olga this is not a problem to solve and forget. It is an ongoing infrastructure challenge. So how do you solve it? “What BGE has taught us is that future large-scale biodiversity genomics efforts – including the European Reference Genome Atlas – will need a living, shared, openly accessible collection of taxon-specific protocols that grows with every new species we tackle.” BGE has laid the groundwork for these issues, but the real work of maintaining and expanding it lies ahead. “The difficult species are not the edge cases. They are the centre of what biodiversity genomics is about.”
More than 20,000 samples have eventually been successfully sequenced, all due to the hard work of Olga and her colleagues. The next step is to make all this hard won data ‘FAIR’: Findable, Accessible, Interoperable and Reusable. Curious? We will share more about this in our next article!
Photo credit: Raffaele de Pascalis / Naturalis Biodiversity Center





