The BGE consortium has undertaken an international effort to sample biological material through fieldwork campaigns and from multiple natural history collections. The consortium incorporates collections and scientists covering a broad spectrum of taxa and expertise. Citizen scientists have also provided crucial input, through their own collected and identified specimens, ‘bioblitz’ events, and carrying out community and eDNA sampling.
All sampling has complied with legal and ethical protocols, with collecting permits in place. All samples – of both individual organisms and communities – were archived in biobanks and associated metadata properly recorded and deposited in public repositories to ensure they are accessible for future use and verification.
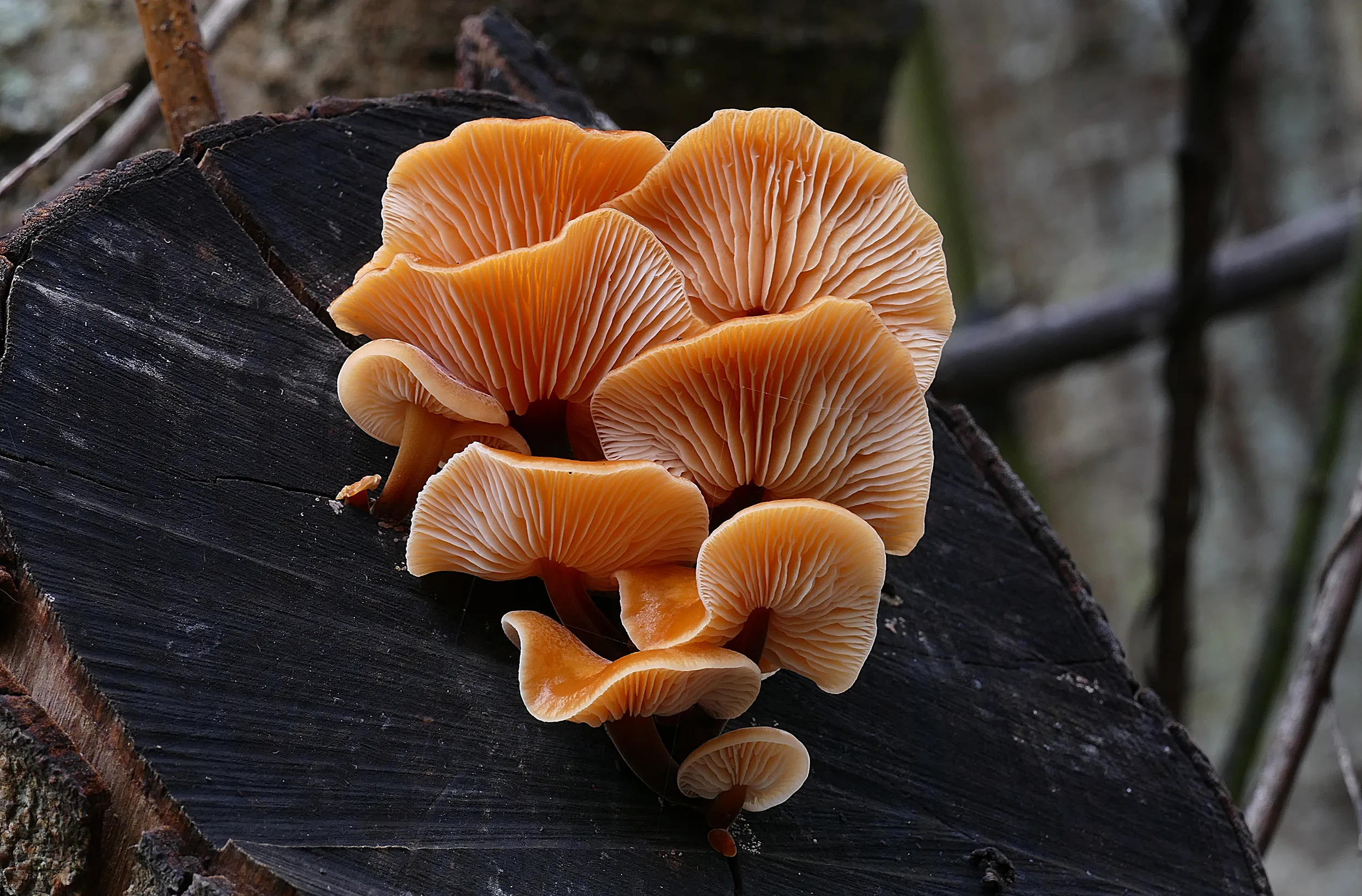
Two and a half years after the project’s launch, the sampling phase for reference genome production has been successfully concluded. A total of 1,113 eukaryotic species were collected and 314 species delivered to sequencing facilities following the established species selection prioritization scheme. The delivered species span 23 countries, representing 16 phyla, 118 orders and 186 families. The three most represented phyla are Arthropoda, Streptophyta and Chordata.
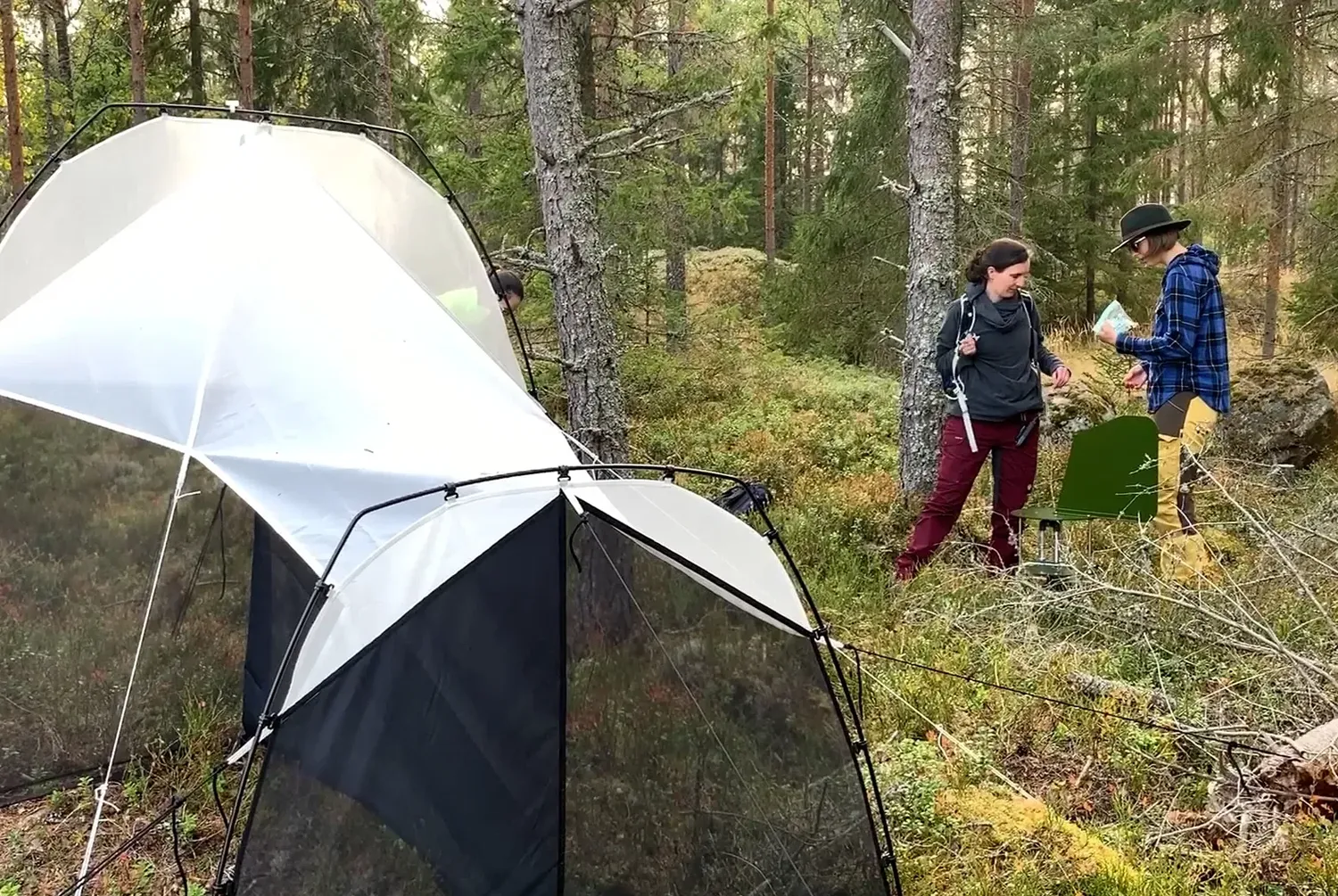
Species were collected using two distinct sampling strategies: a community-driven sampling effort conducted across European countries, and sampling biodiversity hotspots. The first strategy involved two open calls to the ERGA community for species nominations encouraging strong community engagement. The second was a coordinated effort focused on targeting the Mediterranean.
To further increase geographic and phylogenetic representation, additional species were collected during Bioblitz events by subcontracted parties, also members of the ERGA community. Most of the delivered species were preserved as morphological vouchers in museum collections in addition to archiving in biobanks.
Overcollection strategies were implemented as a proof of concept for scalability, ensuring sufficient sample availability. Oversampled material that was not delivered for genome sequencing was stored at the Natural History Museum of Madrid and made available for potential use in future relevant initiatives, fostering collaboration and supporting further research efforts. Considering BGE’s genome prioritizing goals that focus on species relevant to agriculture, fisheries, key ecosystem processes, as well as endemic, threatened species, pests and disease vectors, our target species list aligned well with the established priorities.
Sampling for the DNA-barcode reference library has taken advantage of the project developed gap list of missing species to direct BGEs efforts towards key groups missing in the Barcode of Life Data Systems (BOLD), including insect pollinators, aquatic (freshwater and marine) invertebrates, fish and plants.
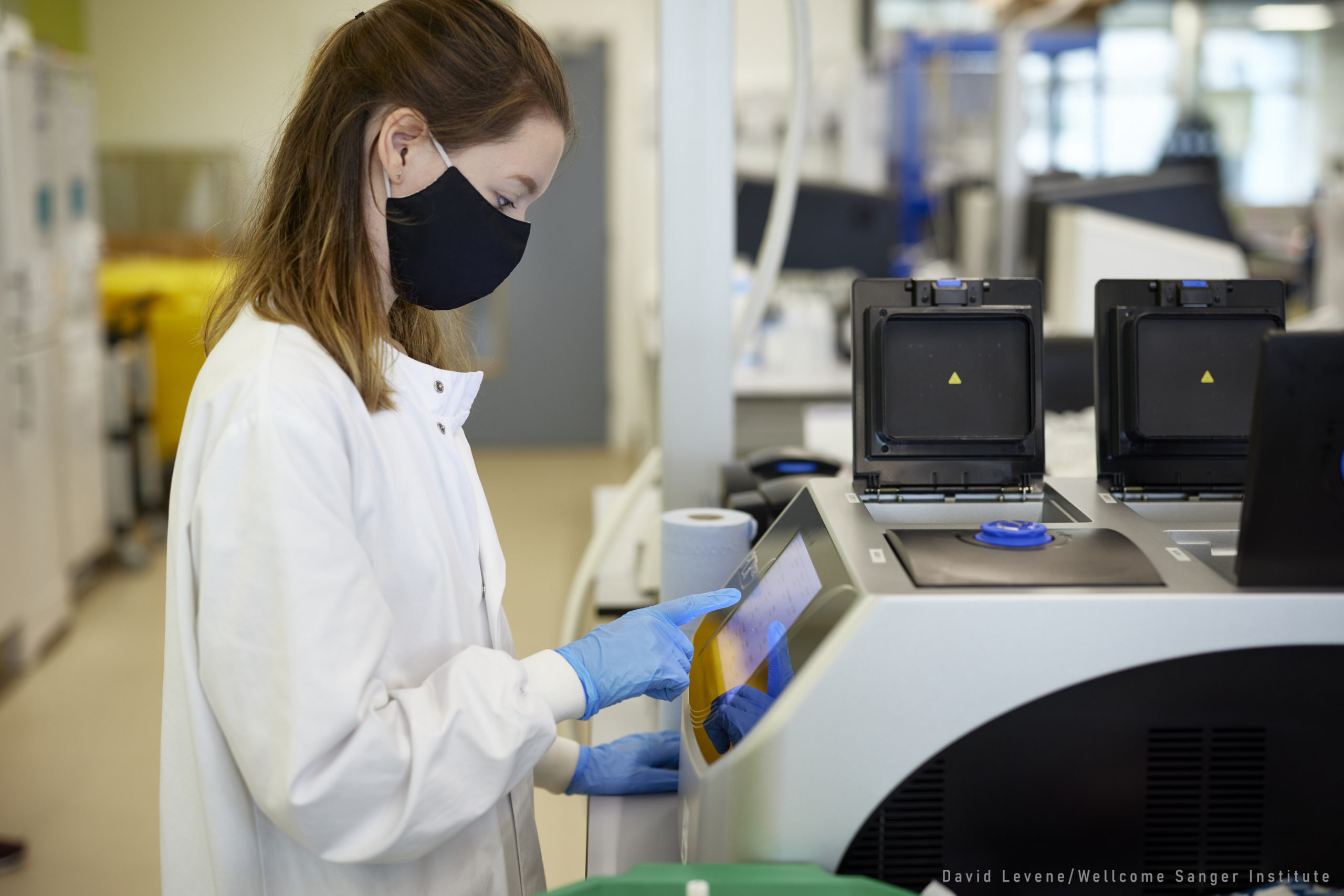
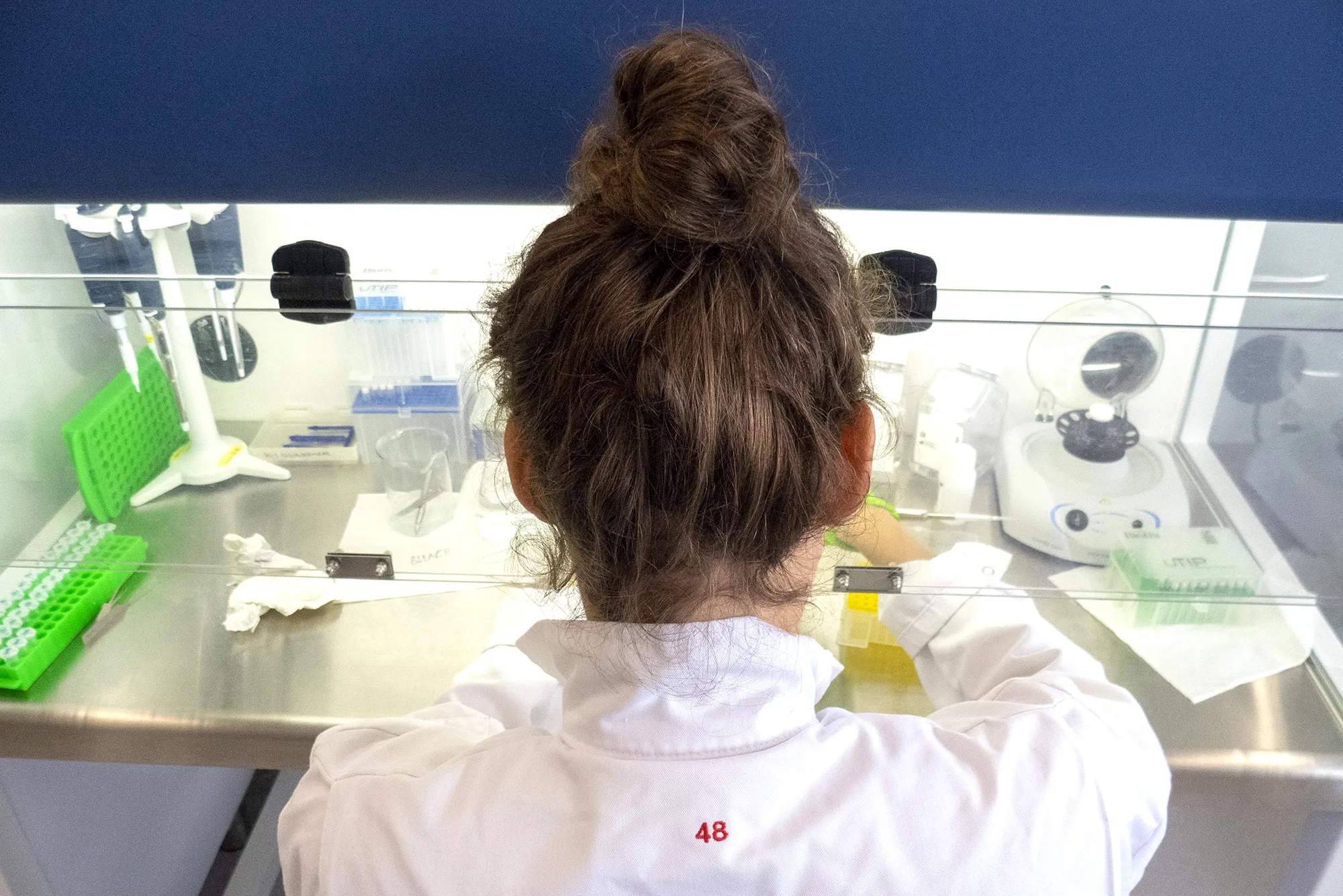
Two and a half years in, more than 42,000 specimens have been sampled with tissues submitted for either amplicon sequencing (if fresh specimens) or genome skimming (if historical specimens). Care has been taken to ensure that vouchers of all species are deposited in natural history collections.
The community sampling focused on several different sample types, organism groups and habitats. Soil was sampled to investigate progress in ecological restoration at targeted sites, water was filtered from European ports and harbours to detect marine invasive species, and insect communities were sampled with Malaise traps to compare species composition along elevation gradients across selected European mountains or pollinator communities in gardens versus agricultural sites.
Several hundred sites have been visited and over 5,500 samples were taken, many hundreds of which were contributed by citizen scientists and other stakeholders.